Deconstructing The Amyloid Fiber
Amyloids

A hallmark of many neurodegenerative diseases, including Parkinson’s and Alzheimer’s, is the abnormal aggregation of proteins into fibrils called amyloids. One possible therapeutic approach would be to reverse this aggregation. For almost two decades, Dr. David Eisenberg, professor of biological chemistry and an investigator of the Howard Hughes Medical Institute , has been trying to visualize and determine structure of these amyloids. Dr. Eisenberg believes that understanding their molecular properties— the actual atomic structures—of these proteins holds the key to therapies that interrupt disease processes.
"Our basic idea is that if we stop fiber formation, we can halt disease," he says. "What we do is learn the structure of the particular fiber associated with each of these diseases. The next step is we design a molecule which will stop formation of the fiber."
In 2005, his lab determined for the first time the atomic-level structure for an amyloid fiber. Seeing the atomic structure provides clues as to why some proteins form amyloids and others do not and reveals potential targets for drugs to inhibit the formation of the proteins. He is now studying several different amyloid proteins, including the abnormal aggregates of the protein tau that are seen in the brains of people with Alzheimer's disease. His lab has identified the atomic structure for two segments of tau that causes it to form fibers. They are now working on designing a molecule —an inhibitor— to stop the fiber formation.
"We have 40 or more amyloid diseases, and there are no effective drugs for any of those diseases," Eisenberg says. "Why is that? It's because Big Pharma has thought that it's impossible to learn the structures of these targets. We've shown it is possible. Knowing that, one can design inhibitors. I believe it's absolutely key to learn the structures of these protein fibers."
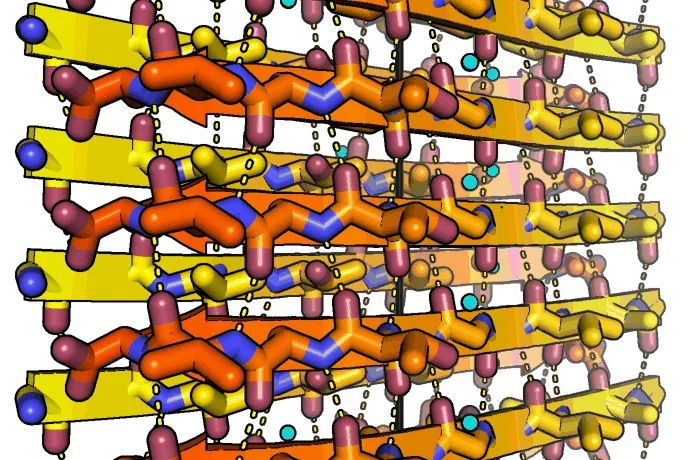
To the left: Amyloid fibrils of human alpha-synuclein, residues 68-78. Alpha-synuclein fibrils are abundant in the cells of Parkinson's disease patients. The peptides self-assembles into sheets through backbone hydrogen bonds (dashed lines). Two sheets mate together tightly into fibrils. The tight fit between sheets lends the fibrils strength. The black line indicates the fibril axis. The structure was determined at 1.4 angstrom resolution using microED. Credit Jose A. Rodriguez and Michael R. Sawaya